Researchers from NTU are utilising NSCC’s supercomputing resources to improve the performance of learning models in molecular data analysis.
AI is expected to play an increasingly important role in the years to come, especially in biological studies. As a major part of AI, neural networks cannot achieve such success without the support of enormous data.
The huge advancements in biological sciences and technologies has led to the accumulation of unprecedented amounts of biomolecular data. For example, in a protein data bank, there are about 150,000 three-dimensional biomolecular structures (Berman, Westbrook et al. 2000). There is an abundance of available biological structures, data analysis methods and models, including data mining, manifold learning and graph or network models. Leveraging the data, topological data analysis (TDA), for example, can potentially provide great promise in the big data era and have become increasingly popular in bioinformatics and computational biology in the past two decades.

Data-driven learning models are among the most important and rapidly evolving areas in chemoinformatics and bioinformatics. Featurization, or feature engineering, is key to the performance of machine learning models in material, chemical, and biological systems. As such, a group of researchers at the School of Physical and Mathematical Sciences at Nanyang Technological University Singapore are using high performance computing to develop a new molecular representation framework, known as persistent spectral (PerSpect), and PerSpect based machine learning (PerSpect ML) for protein-ligand binding affinity prediction. The proposed PerSpect theory provides a powerful feature engineering framework. PerSpect ML models demonstrate great potential to significantly improve the performance of learning models in molecular data analysis.
High performance computing plays a pivotal role in the team’s daily work. The project needs to process large databases which contains thousands of entries, the databases needs to be divided into several pieces and parallel computing has to be employed to treat each part. Additionally, some algorithms are time/memory-consuming and computing resources with multiple cores and large memory are needed to run them.
To find out more about the NSCC’s HPC resources and how you can tap on them, please contact [email protected].
NSCC NewsBytes May 2021
Other Case Studies
Advancing Drug Discovery Research using NSCC HPC resources
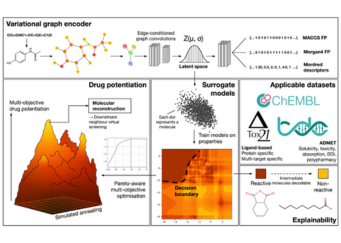
Researchers from Nanyang Technological University (NTU) are applying variational graph encoders as an effective generalist algorithm in computer-aided drug design (CADD)....
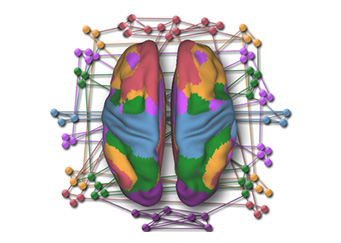
Gaining Deeper Insights into Mental Disorders through Brain Imaging and High-Performance Computing
Researchers from NUS are leveraging supercomputing to develop better strategies for prevention and treatment to mitigate the impact of mental illness. The human brain is a marvel...
Using Digital Twin Technology to Optimise the Industrial 3D Printing Process
Researchers from the Institute of High-Performance Computing (IHPC) are utilizing supercomputers to create a digital twin that furnishes users with comprehensive information...