Researchers at A*STAR are tapping HPC resources to gain a deeper understanding of the dengue virus in order to design improved therapeutic strategies.
Dengue virus (DENV) is a member of the flavivirus genus and infects hundreds of millions of people worldwide each year. A Flavivirus is like an onion; the outermost layer is made up of “envelope proteins” embedded in a lipid membrane, while the inner core contains capsid proteins in complex with an RNA genome.
Experimental structures of viruses correspond to single “snapshots” at a given physiological condition but do not always reflect their dynamic nature or important conformational changes related to binding of antibodies or drugs. For example, consecutive infections by different serotypes of DENV can lead to “antibody dependent enhancement” (ADE) in which antibodies from the first infection bind to the infecting virus particle but do not neutralize it – this can result in a severe case of dengue.
 At present, there are no approved drugs or highly effective and safe vaccines for DENV. Therefore, a team of researchers from Peter J. Bond group at the Bioinformatics Institute (BII) at A*STAR, in collaboration with multiple experimental groups in Singapore, are leveraging on NSCC’s supercomputing resources to get a detailed understanding of the structural and dynamical properties of DENV.
At present, there are no approved drugs or highly effective and safe vaccines for DENV. Therefore, a team of researchers from Peter J. Bond group at the Bioinformatics Institute (BII) at A*STAR, in collaboration with multiple experimental groups in Singapore, are leveraging on NSCC’s supercomputing resources to get a detailed understanding of the structural and dynamical properties of DENV.
The team recently explored the molecular details of ADE and proposed a new mechanism by combining computational methods with experimental snapshots. The team used a “virtual microscope” approach based on molecular dynamics simulations, in order to relate static snapshots of virus-antibody complexes to their dynamics. They particularly focused on the DENV envelope proteins since these
are crucial for the virus to attach to the host cell during infection and are also recognized by antibodies during a host immune response.
Through probing virus dynamics at the molecular level, an improved understanding of the mechanisms of disease and antibody binding can be achieved and the team hopes to ultimately work towards improved therapeutic strategies.
To find out more about the NSCC’s HPC resources and how you can tap on them, please contact [email protected].
NSCC NewsBytes January 2022
Other Case Studies
Advancing Drug Discovery Research using NSCC HPC resources
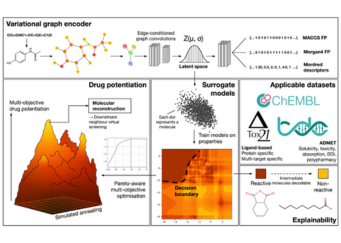
Researchers from Nanyang Technological University (NTU) are applying variational graph encoders as an effective generalist algorithm in computer-aided drug design (CADD)....
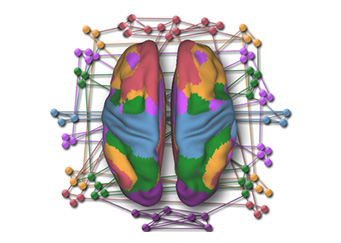
Gaining Deeper Insights into Mental Disorders through Brain Imaging and High-Performance Computing
Researchers from NUS are leveraging supercomputing to develop better strategies for prevention and treatment to mitigate the impact of mental illness. The human brain is a marvel...
Using Digital Twin Technology to Optimise the Industrial 3D Printing Process
Researchers from the Institute of High-Performance Computing (IHPC) are utilizing supercomputers to create a digital twin that furnishes users with comprehensive information...